Single-molecule phenotyping coupled with in situ single-molecule DNA sequencing
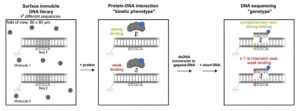
DNA sequencing is a key method that has had a huge impact on diagnostics, genomics and functional analysis. Although many single-molecule sequencing methods exist, there is currently no proficient way to connect the functional properties of a single DNA molecule with its sequence.
Researchers at the University of Oxford have developed a single-molecule sequencing method capable of connecting the functionality (reactions or interactions) of a single molecule with its sequence. This novel method is expected to support screening libraries of candidate therapeutic or agrichemical targets.
Applications: Diagnostics, FRET, SELEX, Aptamer Discovery
Oxford researchers have developed a novel single-molecule sequencing method capable of connecting the functionality of a single nucleic acid molecule with its sequence.
| Features | Benefits |
|
|
|
|
|
|
|
|
Patented & available for:
- Licensing
- Co-development
- Consulting
about this technology

